
Preprocessing
In this stage, images are processed for motion correction, denoising, smoothing, modulation, and alignment between modalities . Each MRI sequence (e.g. T1-weighted, resting) can be classified by a certain MRI modality as shown in the table below.
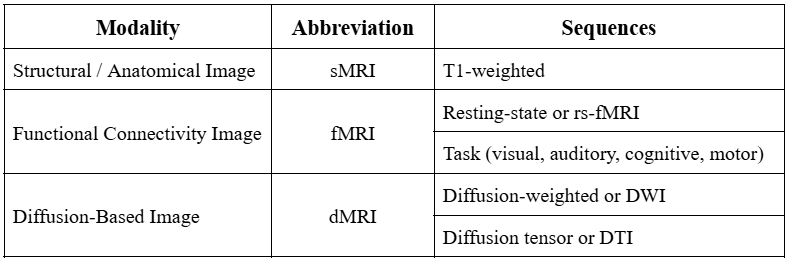
Structural MRI (sMRI) are 3D objects that are used to identify tissue types present in the image. Functional MRI (fMRI) are 4D objects that focus more on the BOLD (blood oxygenation level dependent) signal to map brain activity. Diffusion MRI (dMRI) are 4D objects that use the directionality of water diffusion to map out white matter tracts. Different modalities require different steps of preprocessing. Below summarizes these steps for each MRI modality.
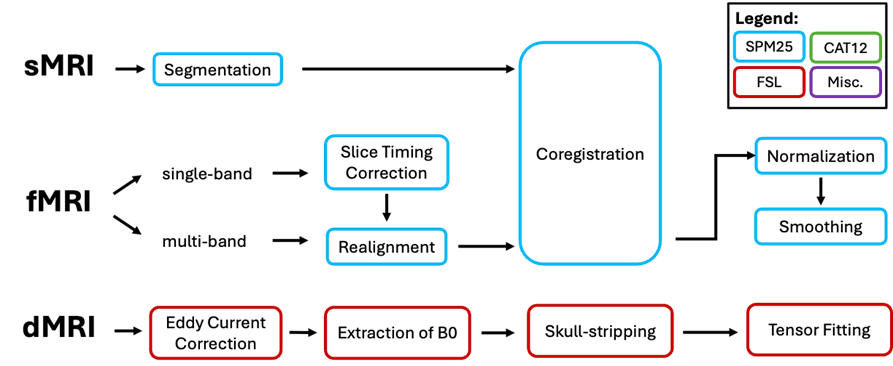
sMRI Preprocessing
Since sMRI stores information about tissue types, it is necessary to segment structural images into separate different structural tissues such as gray matter, white matter, cerebrospinal fluid (CSF), blood, bones, etc. The segmented images are used later for fMRI coregistration to a common space
fMRI Preprocessing
Functional images can be acquired in two frequency bands: single-band, or multi-band. Single band acquisition uses longer repetition time (TR) and thus takes longer duration of acquisition. However, older MRI scanners may not have access to multi-band sequencing. For single-band fMRI, it is necessary to correct motion artifacts in time according to a reference slice. Then, the images are realigned to correct motion artifacts in space. The functional images are then coregistered with sMRI and normalized to the Montreal Neurological Institute (MNI) Space for better mapping of brain regions. Smoothing of functional images increases signal-to-noise ratio (SNR) by taking the weighted average of the neighboring pixels
dMRI Preprocessing
Diffusion-based images first undergo eddy current correction. Eddy currents are void-like structures in the images that are induced by the external magnetic field during acquisition. These are removed to improve accuracy of the tract-based analysis. Then, the first slice is extracted assuming that this slice is acquired when the magnetic field strength is still zero, The extracted slice is used as a mask for skullstripping, to remove the bones and facial structures. Finally, a tensor model is fitted utilizing the given eigenvectors of directionality
PET/CT Preprocessing
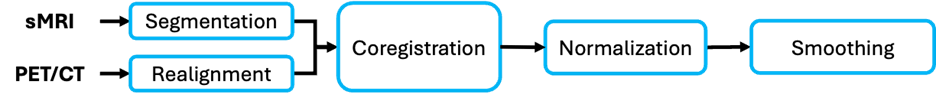
PET images are 3D objects that store information about metabolic activity of different regions of the brain. PET images can be treated similar to functional images except that the preprocessing does not involve slice timing correction. Below are the steps in preprocessing PET images


Imaging Data and Analysis
Clinical and Neuropsychological Assessment
Structural MRI
Functional MRI
DTI
FDG-PET
Neuromap-PH
info@neuromap.ph
